Using the BIDS Data Standard with RT-Cloud
Note: Some of this documentation is taken from Polcyn, S. (2021) “Efficient Data Structures for Integrating the Brain Imaging Data Structure with RT-Cloud, a Real-Time fMRI Cloud Platform” [Unpublished senior thesis].
BIDS Introduction
BIDS is the leading data standard for neuroscience data and is supported by a wide variety of data formatting and analysis tools. It is the standard used by OpenNeuro which is a large and growing repository of neuroscience datasets. In addition there are a large set of BIDS Apps, which are container-based applications with a standardized interface that work on BIDS-formatted data. The BIDS Validator is an automated and comprehensive validation tool that analyzes datasets and identifies BIDS compliance issues.
The BIDS Archive
The BIDS standard defines the on-disk layout and format of datasets to form a BIDS archive. A BIDS archive is a collection of brain activity image and metadata files for one study, which may comprise multiple subjects across multiple days. While an in-depth understanding of the BIDS standard can be obtained from the full standard, viewable online at https://bids-specification.readthedocs.io/en/stable/, a few key details are as follows:
Brain imaging data is stored in the Neuroimaging Informatics Technology Initative (NIfTI) format. NIfTI is a binary file format that starts with a header holding basic information about the brain data contained in the file. The header is followed by the raw brain data. A NIfTI volume’s data typically has 4 dimensions, x, y, z and t (time), so a NIfTI file can be thought of as containing a sequence of t, 3-D images, each of which has dimensions
x * y * z.Metadata is stored in files separate from the image data. Unlike the DICOM image format, the NIfTI image format doesn’t store much metadata about the image it contains. Accordingly, in the BIDS data format, the majority of the metadata about the image and the conditions under which it was collected is stored in separate files, typically in the JavaScript Object Notation (JSON) or Tab-Separated Value (TSV) format.
Files are named using BIDS entities. The name of a file in a BIDS archive follows a standard format, and it is composed of a set of ‘entities’ (like ‘sub’ or ‘run’, corresponding to ‘subject’ and ‘run’, respectively) that signify what the data in the file corresponds to. For example, the filename “sub-01_task-language_run-1_bold.nii” has 4 BIDS entities, separated by underscores (‘_’). The 4 entities and their corresponding values are:
‘sub’: 01 (this file has data for the subject with ID 01)
‘run’: 1 (this file has data from the 1st run)
‘task’: language (this file has data from the ‘language’ task)
‘bold’: No value (The presence of the entity is enough to state the file holds fMRI brain-oxygen-level-dependent (BOLD) data)
In summary, a BIDS archive is a collection of image and metadata files, all named using BIDS entities that correspond to the conditions under which the data or metadata was collected.
BIDS Apps
BIDS Apps are containerized applications that operate on BIDS datasets and provide a consistent command-line interface to users. Since each app operates on a BIDS archive, a full analysis pipeline can potentially be created from independent BIDS App containers, so components can be easily added, removed, or modified as needs evolve over time.
Why Use BIDS with RT-Cloud
BIDS Apps
Using BIDS with RT-Cloud connects you to the BIDS Apps ecosystem, so you can integrate existing and future BIDS Apps with your real-time fMRI analysis pipeline, minimizing time spent on setting up computational infrastructure.
Benefits of BIDS Data Standardization
Storing data in a standardized format brings a host of benefits, the following of which were adapted from here.
One major benefit is you and all lab members or clinical team members, once having learned the standard, know immediately how to navigate both new and old datasets. Without a standardized format, different team members may format their data in different ways, forcing you to waste time learning a myriad of data formats and creating significant problems when a team member leaves the organization and can no longer explain to new or existing team members how their dataset format works. Additionally, external collaborators at other institutions can easily work on your dataset if everyone uses the same standard.
Another major benefit is future software packages are likely to grow around this standard. Thus, you can use any of a wide variety of software packages with your new and existing BIDS datasets that conform to the standard and not spend time learning additional software-specific formats or be locked-in to a particular software package.
Finally, if you are required to publish your datasets as a condition of manuscript publication, having data in a standardized format from the beginning enables a seamless upload and review process.
Adapting BIDS for use in Real-Time fMRI Experiments
Real-time fMRI experiments involve processing image data as it arrives from the scanner and providing immediate subject feedback. In essence, rt-fMRI is a streaming model, whereas BIDS is a data-at-rest standard. To adopt BIDS for rt-fMRI we introduce a new idea, the BIDS Incremental.
A BIDS Incremental packages one brain volume into its own BIDS archive. Thus, we can use this to send a stream of very small BIDS archives (i.e., BIDS Incrementals) for processing. This allows the processing to be done by any application that can ingest BIDS data, such as BIDS-Apps.
How to Incorporate BIDS into your RT-Cloud project
There are three primary classes to use to leverage BIDS in your RT-Cloud project: BIDS Incremental, BIDS Run, and BIDS Archive.
BIDS Incremental is a single-image data structure, encapsulating a single-volume BIDS Archive.
BIDS Run is a data structure that efficiently stores a full run’s worth of BIDS Incrementals in-memory and in a deduplicated fashion. It supports appending BIDS Incrementals to a scanning run and retrieving BIDS Incrementals that have already been added.
BIDS Archive is a data structure that provides an API for interacting with on-disk BIDS archives and enables efficient movement between the BIDS Run streaming data structure and the on-disk BIDS archive.
Below is a sample of how your project can receive real-time scanner data in BIDS-incremental format. This assumes you are running the scannerDataService in the control room. This example communicates with the scannerDataService via the clientInterface.bidsInterface. A data stream is initialized, giving the scanner directory that the DICOMs will arrive in and the DICOM filename pattern to watch for.
from rtCommon.clientInterface import ClientInterface
# connect to the remote data service (via projectServer on localhost)
clientInterfaces = ClientInterface()
bidsInterface = clientInterfaces.bidsInterface
# specify the BIDS entities for the run being done
entities = {'subject': cfg.subjectName,
'run': cfg.runNum[0],
'suffix': 'bold',
'datatype': 'func',
}
# initialize the stream which will watch for DICOMs created at the scanner
# and then convert them to BIDS-incrementals and stream them to this script.
streamId = bidsInterface.initDicomBidsStream(cfg.dicomDir,
cfg.dicomScanNamePattern,
cfg.minExpectedDicomSize,
**entities)
# loop over the expected number of DICOMs per run
for idx in range(scansPerRun):
bidsIncremental = bidsInterface.getIncremental(streamId, idx)
imageData = bidsIncremental.imageData
avg_niftiData = numpy.mean(imageData)
if cfg.writeBidsArchive is True:
# See openNeuroClient project under 'projects' directory for more
# information on accumulating a BIDS archive from a stream of incrementals.
newRun.appendIncremental(bidsIncremental)
Below is a simple example that shows the interactions between the various classes.
archive = BidsArchive('/tmp/bidsDataset')
print('Subjects:', archive.getSubjects(), 'Runs:', archive.getRuns())
# Query the run using BIDS Entities (see the tutorial for a deeper introduction)
run = archive.getBidsRun(subject='01', run=1, datatype='func')
newRun = BidsRun()
meanActivationValues = []
for i in range(run.numIncrementals()):
incremental = run.getIncremental(i)
meanActivationValues.append(np.mean(incremental.imageData))
newRun.appendIncremental(incremental)
newArchive = BidsArchive('/tmp/newBidsDataset')
newArchive.appendBidsRun(newRun)
An overview of how these classes all fit together for sending data from the MRI
scanner to a BIDS Archive is shown here:
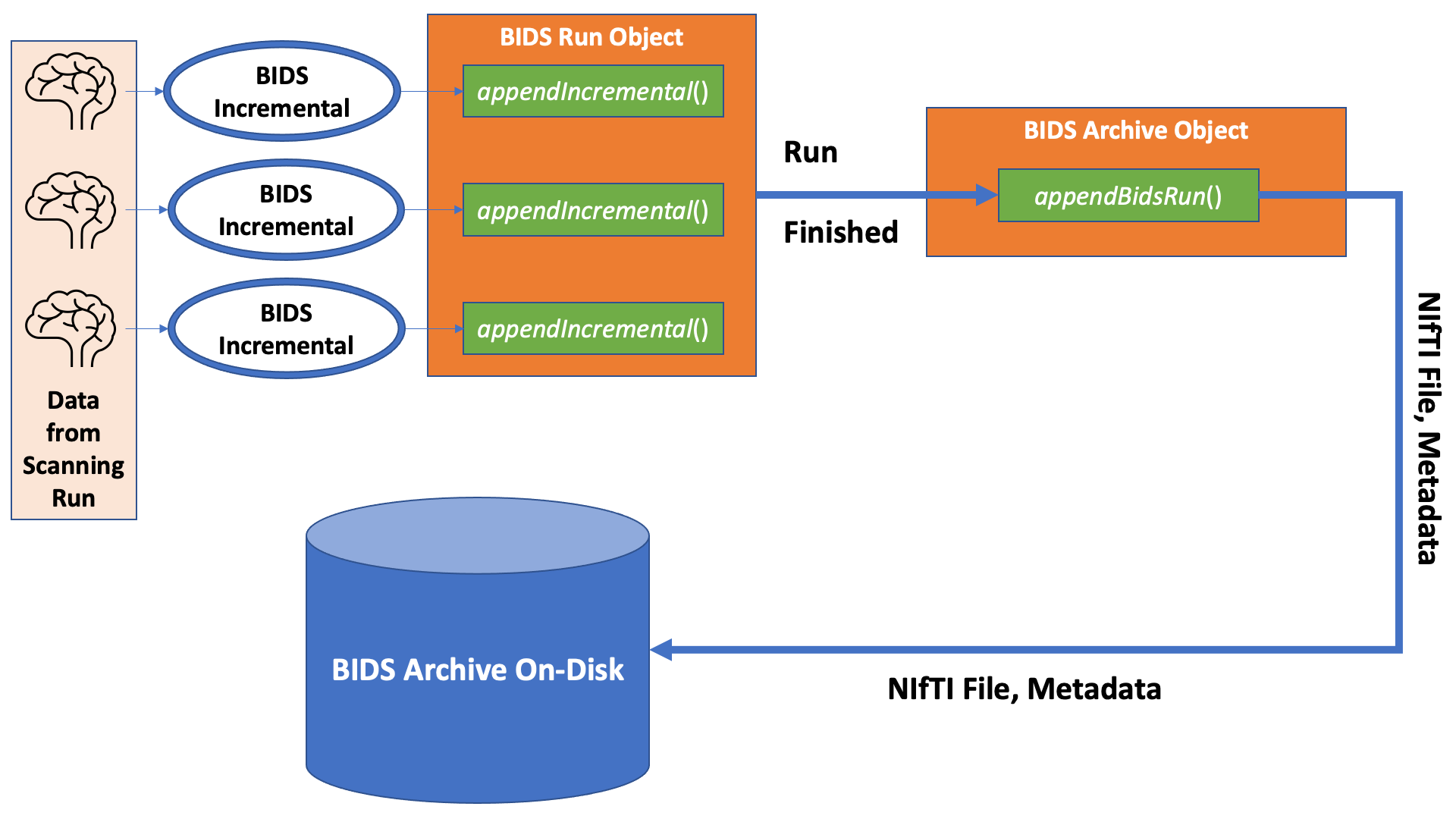 Retrieving BIDS data from an archive is simply the reverse of that diagram.
Retrieving BIDS data from an archive is simply the reverse of that diagram.
For a more in-depth introduction to the various classes and how to use them, check out the bids_tutorial Jupyter notebook tutorials/bids_tutorial.ipynb.
Replaying Data from OpenNeuro
One goal of this project is to facilitate collaboration and sharing of code and data. To this end we introduce an OpenNeuro module which can access and stream data from the OpenNeuro.org data repository. In essence this is a ‘NetFlix’ type service for fMRI datasets. Researchers can replay datasets through their processing pipelines to try new models, reproduce results or test and debug experiments.
An example of streaming OpenNeuro data can be seen in the projects/openNeuroClient sample project. The key snippets of code are shown below.
# OpenNeuro accession number for a dataset
dsAccession = 'ds002338'
# The subject and run number to replay
entities = {'subject': 'xp201', 'run': 1}
# Initialize the data stream
streamId = bidsInterface.initOpenNeuroStream(dsAccession, **entities)
numVols = bidsInterface.getNumVolumes(streamId)
# Retrieve and process each volume as a BIDS-Incremental
for idx in range(numVols):
bidsIncremental = bidsInterface.getIncremental(streamId, idx)
imageData = bidsIncremental.imageData